
The AP Biology exam for 2017 is set for the morning of Monday May 8th. So it’s that time of the year again to begin reviewing past concepts and doing practice exams.
The free response questions on the past four exams didn’t touch so much on a very important concept, the tools and techniques of biotechnology. So, in this post we’ll review some of the major tools and techniques while going over Question 4 from the 2007 AP Biology Exam. On average students scored 2.52/10 points. This post will show students how they could have maximized their score and give a review of important terms.
The Question
A bacterial plasmid is 100kb in length. The plasmid DNA was digested to completion with two restriction enzymes in three separate treatments: EcoRI, HaeIII, and EcoRI + HaeIII (double digest). The fragments were then separated with electrophoresis, as shown.
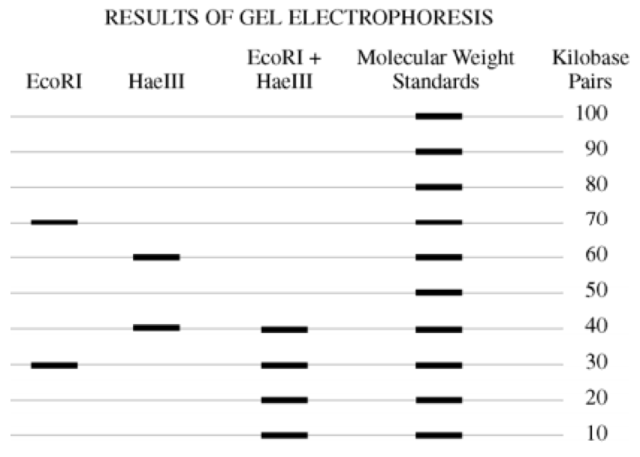
a) Using the circle provided, construct a labeled diagram of the restriction map of the plasmid. Explain how you developed your map.
b) Describe how:
- Recombinant DNA technology could be used to insert a gene of interest into a bacterium
- Recombinant bacteria could be identified
- Expression of the gene of interest could be ensured
c) Discuss how a specific genetically modified organism might provide a benefit for humans and at the same time pose a threat to a population or ecosystem.

The Solution
In order to answer this question you first had to know what they were asking for, so let’s review some terms.
Plasmid
A plasmid is a circular double-stranded DNA molecule usually found in bacteria that is capable of replicating independently of the cell’s chromosomal DNA. Usually plasmids allow the bacteria to develop some advantage such as antibiotic resistance. A bacterial cell can have more than one plasmid, will express the genes on those plasmids and will replicate each plasmid when it divides. In this way, each daughter cell will get a copy of the plasmid. Plasmids are often used for cloning purposes since genes of interest (fragments of DNA) can be inserted into plasmids and introduced into the bacteria by bacterial transformation. In bacterial transformation, competent bacteria where the cell wall has been made permeable to genetic material can take up a foreign plasmid. The bacteria that have taken up the plasmid can be selected for usually by growing the bacteria in a certain antibiotic to which the plasmid DNA allows resistance. In this way, as the bacteria divide the plasmid and thus the genes on them will be amplified.
Restriction enzymes
These are enzymes that cut DNA at specific recognition sites that are usually 4 to 8 base pairs in length. The sites are usually also palindromic, meaning they read the same forwards and backward. The restriction enzymes will also produce “sticky” or “blunt” ends. These ends can be used to insert the gene of interest into a plasmid by ligation.
Ex. Sticky end where bases are left unpaired after a cut
 Ex. Blunt end where all bases are paired after a cut
Ex. Blunt end where all bases are paired after a cut

Electrophoresis
Gel electrophoresis separates molecules of DNA on the basis of their rate of movement through an agarose gel in an electric field. Remember that DNA is negatively charged due to its phosphate backbone and will flow from cathode (-) to anode (+). Smaller fragments of DNA will travel faster and thus be farther away from the starting wells. Similarly, proteins can also be separated using electrophoresis.
Restriction mapping
Using restriction enzymes and gel electrophoresis the restriction sites on a segment of DNA, usually a plasmid, can be mapped. This is useful when trying to identify whether a gene of interest was correctly inserted into the plasmid.
Part (a)
Using the circle provided you were to draw and label a restriction map of the resulting gel electrophoresis as well as explain how you came up with the map. The restriction enzyme EcoRI resulted in two bands of DNA, one that was 70kb and another that was 30kb. The whole plasmid is 100kb so from this information you can deduce that there are two sites where EcoRI cuts the DNA, since 70 + 30 = 100. So place two EcoRI sites on your circle.
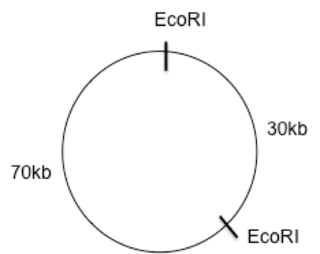
Now we look at the fragments that are created by HaeIII. A 60kb and a 40kb fragment which equals 100kb as well. So we know there are only two sites where HaeIII cuts as well. Hold off on putting the HaeIII sites. The last restriction digest was a double digest using both EcoRI and HaeIII. Since both EcoRI and HaeIII result is two cuts each using a double digest should result in 4 fragments. In this case those fragments are 40, 30, 20, and 10kb. Using trial and error place the HaeIII cut sites so that the distance between the EcoRI sites doesn’t change and so that you get four cuts of the sizes 40, 30, 20, and 10kb. I would start by splitting the 70kb into two cuts one of 40kb and one of 30kb. And split the 30kb into one 20kb and one 10kb as shown below.
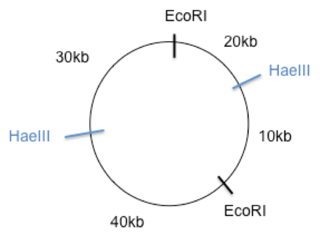 However, you aren’t done. You need to make sure that the HaeIII sites are 60kb from each other one way and 40kb the other. In our map above this is not the case so you should rearrange the numbers.
However, you aren’t done. You need to make sure that the HaeIII sites are 60kb from each other one way and 40kb the other. In our map above this is not the case so you should rearrange the numbers.
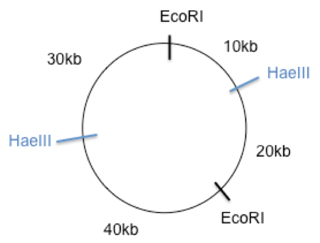 The resulting map should look something like this. Where EcoRI cuts at two sites that result in 70kb and 30kb fragment and where HaeIII results in a 60kb and 40kb fragment. In your explanation you should describe the process of trial and error that you used to come to this conclusion. DO NOT waste time providing information that was not asked for as this wastes valuable time. So there is no need to explain gel electrophoresis!
The resulting map should look something like this. Where EcoRI cuts at two sites that result in 70kb and 30kb fragment and where HaeIII results in a 60kb and 40kb fragment. In your explanation you should describe the process of trial and error that you used to come to this conclusion. DO NOT waste time providing information that was not asked for as this wastes valuable time. So there is no need to explain gel electrophoresis!
Part (b)
This section asks you to describe how a gene of interest is inserted into a bacterium, how recombinant bacteria (bacterially that have been genetically modified) can be identified and how expression of that gene of interest can be ensured. First, the gene of interest needs to be cut from the source, which can be accomplished using a restriction enzyme. This same restriction enzyme should be used to cut the plasmid that the gene will be inserted into to ensure compatible ends. The gene of interest and plasmid then need to be combined so that the complementary ends can match up and join together. Once the gene of interest is inserted into the plasmid, this plasmid needs to be incubated with competent cells so that these cells can take this plasmid up. To identify whether the bacteria has taken up the plasmid the bacteria can be grown on antibiotic plates as long as the plasmid has a gene for antibiotic resistance. To ensure that the gene of interest is being expressed the gene of interest can be inserted downstream of a promoter that can be induced when giving certain nutrients such as lactose. Remember that the promoter is the region on DNA that initiates transcription. Make sure you understand the process of gene cloning because it’s bound to come up on the exam! Here is a link to a great overview.
Part (c)
Genetically modified organisms are the results of genes being artificially inserted into the genome of organism. They are used all across science to produce new forms of drugs, vaccines, and even to produce food. Bacteria have been modified to produce insulin to help patients with diabetes, model organisms are genetically modified in research settings to test the affects of a certain gene product, and crops can be modified to produce a certain color, smell, or prevent against plant disease. Genetically modified organisms provide a benefit for humans in many ways especially when it comes to growing large amounts of food. Crops can be modified to become resistant to certain bacteria that threaten the yield thus increasing the amount of food available for consumption. At the same time this poses a risk because the bacteria may become resistant to the antibiotic the crop is producing and cause harm in the long term. For this part it was important to be as specific as possible and describe your ideas in a scientific manner.
The AP Biology exam is meant to be challenging but knowing how commonly used tools and methods are used is essential! Make sure to answer each part of the question completely and label all your answers to maximize your score!
Comments